Simple Booster
Simplified model of the Fermilab Booster with 24 cells and 22 RF cavities. The initial total voltage at injection is 200 KV and the Booster operated a harmonic number of 84.
The Booster is a 474.202 m long rapid cycling synchrotron built in 1969-71 designed to rapidly accelerate protons from a kinetic energy of 0.8 GeV to 8 GeV although this example just runs at the injection energy. The architecture of the accelerator is a modified FODO design and is usually described as “FOFDOOD” which results from splitting the F and D magnets in two, then elongating the space between the two D magnets. The F and D magnets combine both bending and focussing(defocussing) functions to economomize on space which is quite limited.
The lattice is read from the file booster_impactx_lattice.py which is
based on the original MAD-X file sbbooster-cooked-rfon.madx.
The matched Twiss parameters determined by both Synergia and MAD-X at entry are:
\(\beta_\mathrm{x} = 33.73645362843065243\) m
\(\alpha_\mathrm{x} = -0.01.298673960026007664\)
\(\beta_\mathrm{y} = 5.252517912567207681\) m
\(\alpha_\mathrm{y} = 0.006089861210659328755\)
\(\mathrm{D}_\mathrm{x} = 3.785167992\) m
\(\mathrm{Dp}_\mathrm{x} = 0.001377568703\)
The initial beam parameters follow the PIP-II Booster beam at injection as described in the Conceptual Design Report. The beam consists of protons at kinetic energy 800 MeV with emittances specified as:
\(\epsilon_{x}\) |
\(16 \pi\) mm-mr normalized 95% |
\(\epsilon_{y}\) |
\(16 \pi\) mm-mr normalized 95% |
\(\epsilon_{L}\) |
0.1 eV-s 97% |
Run
This example can only be run with a python script:
Python script:
python3 run_simple_booster.py
For MPI-parallel runs, prefix these lines with mpiexec -n 4 ... or srun -n 4 ..., depending on the system.
examples/simple_booster/run_simple_booster.py. The file booster_impactx_lattice.py from the same directory is also required.#!/usr/bin/env python3
#
# Copyright 2022-2023 ImpactX contributors
# Authors: Eric G. Stern, Axel Huebl, Chad Mitchell
# License: BSD-3-Clause-LBNL
#
# -*- coding: utf-8 -*-
from booster_impactx_lattice import get_lattice
from scipy.constants import c, eV, m_p, pi
from impactx import ImpactX, distribution, elements, twiss
mp_mev = 1.0e-6 * m_p * c**2 / eV
total_Booster_charge = 6.7e12 # PIP-II full Booster
active_buckets = 81 # 81 out of 84 buckets full
harmonic_number = 84
turns = 2
emit_x = 16.0e-6 # normalized 95% emit
emit_y = 16.0e-6
emit_eV_s = 0.1 # longitudinal emittance 97% eV-s
#
# these lattice functions are calculated with Synergia3
# from the sbbooster-cooked.madx file.
alpha_x = -1.298673960026007664e-02
beta_x = 3.373645362843065243e01
alpha_y = 6.089861210659328755e-03
beta_y = 5.252517912567207681e00
disp_x = 3.187407765856291153e00
disp_px = 1.136005067625678322e-03
# s betax alphax psix dispx dprimex betay alphay psiy dispy dprimey
# 4.742027519999999186e+02 3.373645362843065243e+01 -1.298673960026007664e-02 4.216017341852963085e+01 3.187407765856291153e+00 1.136005067625678322e-03 5.252517912567207681e+00 6.089861210659328755e-03 4.278849194102631515e+01 0.000000000000000000e+00 0.000000000000000000e+00
sim = ImpactX()
# set numerical parameters and IO control
sim.space_charge = False
# sim.diagnostics = False # benchmarking
sim.slice_step_diagnostics = True
# domain decomposition & space charge mesh
sim.init_grids()
# set the bucket length so that particles can be phase wrapped
booster_lattice = get_lattice()
total_s = sum(element.ds for element in booster_lattice)
print(f"Read Booster lattice, {len(booster_lattice)} elements, len = {total_s}")
bucket_length = total_s / harmonic_number
print("bucket_length: ", bucket_length)
sim.particle_container().set_bucket_length(bucket_length)
sim.particle_bc = "periodic"
# load a 800 MeV proton beam
kin_energy_MeV = 800.0 # reference energy 800 MeV
bunch_charge_C = eV * total_Booster_charge / active_buckets # used with space charge
npart = 10000 # number of macro particles
# reference particle
ref = sim.particle_container().ref_particle()
ref.set_species("proton").set_kin_energy_MeV(kin_energy_MeV)
distr = distribution.Gaussian(
**twiss(
beta_x=beta_x,
alpha_x=alpha_x,
emitt_x=emit_x / (ref.beta_gamma * 6), # normalized 95% emit -> geometric
beta_y=beta_y,
alpha_y=alpha_y,
emitt_y=emit_y / (ref.beta_gamma * 6), # normalized 95% emit -> geometric
emitt_t=emit_eV_s
* 1.0e-6
* c
/ (ref.mass_MeV * ref.beta_gamma * 6 * pi), # 97% emit eV-s -> RMS emit
beta_t=1258.0, # seems to go from 1158 close to the center to
# 1258 at about 1.25m
# dispersion from Synergia so it needs conversion from dp/p to dE/p
dispersion_x=disp_x / ref.beta,
dispersion_px=disp_px / ref.beta,
)
)
sim.add_particles(bunch_charge_C, distr, npart)
# add beam diagnostics
monitor = elements.BeamMonitor("monitor", backend="h5")
zerodrift = elements.Drift(ds=0.0)
# initial 0 length drift is to force initial phase wrapping
# run a lattice with a single monitor to capture the initial beam.
sim.lattice.clear()
sim.lattice.append(zerodrift)
sim.lattice.append(monitor)
sim.periods = 1
sim.track_particles()
# clear the lattice and load the Booster lattice
sim.lattice.clear()
sim.lattice.extend(get_lattice())
sim.lattice.append(monitor)
# run simulation
sim.periods = turns
sim.track_particles()
# clean shutdown
sim.finalize()
"""
Fermilab Booster lattice translated from the MAD-X description in
``sbbooster-cooked-rfon.madx`` to ImpactX elements.
The ring has 24 super-periods (cells). The first 13 cells use a plain ``dlong``
drift in the middle; the remaining 11 cells (14-24) replace that drift with an
RF station (two ``ShortRF`` cavities separated by drifts). All cells are
otherwise identical; only the F/D bend element names carry the cell index
suffix (e.g. ``fmagu01`` ... ``fmagu24``).
"""
from impactx import elements
# ---------------------------------------------------------------------------
# shared parameters for F and D combined-function bends
# ---------------------------------------------------------------------------
_F_EDGE = dict(
K0=1.6449340668482264,
K1=0.0,
K2=0.0,
K3=0.16666666666666666,
K4=0.0,
K5=0.0,
K6=0.0,
R=1.0,
dx=0.0,
dy=0.0,
g=0.0,
model="linear",
psi=0.0353710910981508,
rc=40.847086,
rotation=0.0,
)
_F_CFBEND = dict(
aperture_x=0.0,
aperture_y=0.0,
ds=2.889612,
dx=0.0,
dy=0.0,
int_order=6,
k_normal=[0.024481550532148122, 0.05410921561, 0.00546850017312],
k_skew=[0.0, 0.0, 0.0],
mapsteps=6,
rotation=0.0,
unit=0,
)
_D_EDGE = dict(
K0=1.6449340668482264,
K1=0.0,
K2=0.0,
K3=0.16666666666666666,
K4=0.0,
K5=0.0,
K6=0.0,
R=1.0,
dx=0.0,
dy=0.0,
g=0.0,
model="linear",
psi=0.03007875592383836,
rc=48.034101,
rotation=0.0,
)
_D_CFBEND = dict(
aperture_x=0.0,
aperture_y=0.0,
ds=2.889612,
dx=0.0,
dy=0.0,
int_order=6,
k_normal=[0.020818543059648396, -0.05738855012, -0.037869227516832],
k_skew=[0.0, 0.0, 0.0],
mapsteps=6,
rotation=0.0,
unit=0,
)
# ---------------------------------------------------------------------------
# small builders for the repetitive elements inside a cell
# ---------------------------------------------------------------------------
def _drift(name, ds):
return elements.ExactDrift(
aperture_x=0.0,
aperture_y=0.0,
ds=ds,
dx=0.0,
dy=0.0,
name=name,
rotation=0.0,
)
def _quad(name, k):
return elements.ExactQuad(
aperture_x=0.0,
aperture_y=0.0,
ds=0.024,
dx=0.0,
dy=0.0,
int_order=6,
k=k,
mapsteps=6,
name=name,
rotation=0.0,
unit=0,
)
def _sextupole(name, k_normal, k_skew):
return elements.ExactMultipole(
aperture_x=0.0,
aperture_y=0.0,
ds=0.024,
dx=0.0,
dy=0.0,
int_order=4,
k_normal=k_normal,
k_skew=k_skew,
mapsteps=5,
name=name,
rotation=0.0,
unit=0,
)
def _bend_station(name, edge, cfbend):
"""[entry edge, CFbend, exit edge] for one combined-function dipole."""
return [
elements.DipEdge(name=f"{name}_usedge", location="entry", **edge),
elements.ExactCFbend(name=name, **cfbend),
elements.DipEdge(name=f"{name}_dsedge", location="exit", **edge),
]
def _rf_station():
"""Two ShortRF cavities with their neighboring drifts. Replaces ``dlong``
in cells 14..24. The total length equals the length of ``dlong`` (5.581 m).
"""
rfc_kwargs = dict(
V=9.688990660720476e-06,
dx=0.0,
dy=0.0,
freq=44704409.98524446,
phase=-90.0,
rotation=0.0,
)
return [
_drift("drifta", 0.21),
_drift("drrf", 1.175),
elements.ShortRF(name="rfc", **rfc_kwargs),
_drift("drrf", 1.175),
_drift("driftb", 0.12),
_drift("drrf", 1.175),
elements.ShortRF(name="rfc", **rfc_kwargs),
_drift("drrf", 1.175),
_drift("dmidls", 0.551),
]
# ---------------------------------------------------------------------------
# one full super-period
# ---------------------------------------------------------------------------
def _cell(i, rf):
"""Build super-period ``i`` (1..24). If ``rf`` is True, substitute the
RF station for the ``dlong`` drift (cells 14..24)."""
suf = f"{i:02d}"
middle = _rf_station() if rf else [_drift("dlong", 5.581)]
return [
# upstream short-straight section (small-aperture correctors + diagnostics)
_drift("sa", 0.176),
_drift("hsxx", 0.024),
_drift("vsxx", 0.024),
_quad("qsxx", 0.020254116087679828),
_drift("bpms", 0.024),
_drift("qssxx", 0.024),
_sextupole("sxsxx", [0.0, 0.0, -0.09001070603982472], [0.0, 0.0, 0.0]),
_drift("sssxx", 0.024),
_drift("sb", 0.256),
# upstream F bend
*_bend_station(f"fmagu{suf}", _F_EDGE, _F_CFBEND),
_drift("mins", 0.5),
# upstream D bend
*_bend_station(f"dmagu{suf}", _D_EDGE, _D_CFBEND),
# middle: plain drift ``dlong`` (cells 1..13) or RF station (cells 14..24)
*middle,
# downstream long-straight section (large-aperture correctors + diagnostics)
_drift("hlxx", 0.024),
_drift("vlxx", 0.024),
_quad("qlxx", -0.0738960996629951),
_drift("bpms", 0.024),
_drift("qlsxx", 0.024),
_sextupole("sxlxx", [0.0, 0.0, 8.591792079114004], [0.0, 0.0, -0.0]),
_drift("sslxx", 0.024),
_drift("drifte", 0.251),
# downstream D bend
*_bend_station(f"dmagd{suf}", _D_EDGE, _D_CFBEND),
_drift("mins", 0.5),
# downstream F bend
*_bend_station(f"fmagd{suf}", _F_EDGE, _F_CFBEND),
_drift("sc", 0.6),
]
def get_lattice():
"""Return the 24-cell Fermilab Booster lattice as a list of ImpactX elements."""
lattice = []
for i in range(1, 25):
lattice.extend(_cell(i, rf=(i >= 14)))
return lattice
examples/simple_booster/sbbooster-cooked-rfon.madx.! Simplified Booster lattice
// Added corrector element qsxx, qlxx values for tunes
// Added corrector element sxsxx, sxlxx values for chromaticities
// From JFO 2022-12-08
// The simplified booster lattice is trivial. I have no madx lattice (you could make one
// very easily) - I just use pyorbit classes directly to instantiate a basic cell; it is then
// replicated 24 times. I took the bending magnets lengths and
// strengths directly from the official MADX lattice file.
// The basic cell is
// d1 fmag d2 dmag d3
// d1, d2, d3 : drifts of lengths 0.6 0.5 and 3.0 m
// fmag: focusing bend L = 2.889612 m
// dmag defocusing bend L = 2.889612 m
// total cell length: 19.758 m
// total ring length = 24*19.758 = 474.20 m
// The length, focusing strengths and curvature radius of the
// magnets are as in the booster MADX file.
// If you entered 1 cell correctly, you should get the periodic solution:
// bx = 33.86 m ax = 0
// by = 5.39m ay =0
// For 24 cells, the raw tunes are nux = 7.017 and nuy = 6.675. You will need to tweak the nominal focusing strengths a bit to avoid resonances.
//--------- Nominal Gradient Magnet Definitions
// EGS
// The apparent cell structure is actually:
// D1, l=0.6;
// FMAGU01;
// D2, l=0.5;
// DMAGU01;
// D3, l=6.0;
// DMAGD01;
// D4, l=0.5;
// FMAGD01;
// DR, l=0.6
! Expand structure to include corrector packages:
! Normal short straight
! drift 0.6 (end of previous cell)
! drift 0.176
! corrector package (short) l=0.168
! drift 0.256
! end of short straight
! non-RF cavity long straight
!
! drift 5.581
! correction package (long) l=0.168
! drift 0.251
!
! long straight with RF cavity
!
! drift 0.21
! rfcavity drift-cavity-drift 2.35
! drift 0.12
! rfcavity drift-cavity-drift 2.35
! 4 drifts total 0.551
! corrector packages (long) 0.168
! drift 0.251
! Corrector package:
!
! HLxx, HKICKER, l=0.024
! VLxx, VKICKER, l=0.024
! Q{S|L}xx QUADRUPOLE, l=0.024 (normal)
! MULTIPOLE Q{S|L}ERR, l=0
! MONITOR, l=0.024
! QS{S|L}xx, QUADRUPOLE, l=0.024 (skew)
! MULTIPOLE QS{S|L}ERR, l=0
! SEXTUPOLE SX{L|S}, l=0.024 (normal)
! SEXTUPOLE SS{L|S}, l=0.024 (skew)
!
ke1 = 0.8; !800 MeV kinetic energy at injection
rhof := 40.847086; ! bending radius of focusing magnet
rhod := 48.034101; ! bending radius of defocusing magnet
blength := 2.889612; ! arc length for both F and D magnets
blengthf := 2.889009499; ! physical length (straight length) for F magnet
blengthd := 2.889176299; ! physical length (straight length) for D magnet
!
! The quad field for the gradient magnet is contained in file " qsdqsf.dat" to be read in before this file !
!
! read from file at time step = 7
qsd := -57.38855012e-3;
qsf := 54.10921561e-3;
! These ssd and ssf strengths come from fitting to 01 Dec 2015 chromaticity data
! and predicts chromaticity much better than using the Drozhdin et al measurements above
ssd := -0.04381647074 + ke1*(0.009150934932+ ke1*(-0.0023900895 + ke1*(0.000318068028 - ke1* 1.6353205e-05)));
ssf := -0.006384940088 + ke1*(0.01967542848 + ke1*( -0.006776746 + ke1*(0.00091367565 - ke1* 4.293705e-05)));
!
! Gradient magnets defined by their physical length aith their bend angle
! being defined by the arc length/radius of curvature
!FMAG: RBEND, L = blengthf , ANGLE = blength/rhof, K1 = qsf , K2 = ssf;
FMAG: SBEND, L = blength , ANGLE = blength/rhof, e1=blength/(2*rhof), e2=blength/(2*rhof), K1 = qsf , K2 = ssf, type=fmag;
DMAG: SBEND, L = blength , ANGLE = blength/rhod, e1=blength/(2*rhod), e2=blength/(2*rhod), K1 = qsd , K2 = ssd, type=dmag;
! drifts in the short straight section
mins: drift, l=0.5, type=shortstraight;
sc: drift, l=0.6, type=shortstraight;
sa: drift, l=0.176, type=shortstraight;
sb: drift, l=0.256, type=shortstraight;
! drifts in the long straight section
dlong: drift, l=5.581, type=longstraight; ! for no-RF cavity cells
drifta: drift, l=0.21, type=longstraight; ! start of RF cavity cell
driftb: drift, l=0.12, type=longstraight;
dmidls: drift, l=0.551, type=longstraight; ! (drift in the middle of the longstraight)
drifte: drift, l=0.251, type=longstraight; ! end of RF cavity cell
drrf: drift, l=2.35/2, type=rfaperture;
total_voltage = 0.2; ! 200 KV
NRF = 22; ! 22 active RF cavities
h = 84;
gamma = 1 + 0.8/pmass;
beta = sqrt(1 - 1/gamma^2);
L = 474.202752; ! circumference of Booster ring
freqhz = h * beta * clight/L;
rfc: rfcavity, l=0, harmon=h, volt=total_voltage/NRF, freq=1.0e-6*freqhz, lag=0, type=rfaperture;
! short corrector package
hsxx: drift, l=0.024, type=shortstraight;
vsxx: drift, l=0.024, type=shortstraight;
qsxx: quadrupole, k1=0.0, l=0.024, k1=0.02025411608767982777, type=shortstraight;
! multipole not included
bpms: drift, l=0.024, type=shortstraight;
qssxx: drift, l=0.024, type=shortstraight; // skew quad
! multipole not included
sxsxx: sextupole, k2=0.0, l=0.024, k2=-0.09001070603982472274, type=shortstraight;
sssxx: drift, l=0.024, type=shortstraight; // skew sextupole
cpshort: line=(hsxx, vsxx, qsxx, bpms, qssxx, sxsxx, sssxx);
! long corrector package
hlxx: drift, l=0.024, type=longstraight;
vlxx: drift, l=0.024, type=longstraight;;
qlxx: quadrupole, k1=0.0, l=0.024, k1=-0.07389609966299509614, type=shortstraight;
! multipole not included
bpml: drift, l=0.024, type=shortstraight;
qlsxx: drift, l=0.024, type=longstraight; // skew quad
! multipole not included
sxlxx: sextupole, k2=0.0, l=0.024, k2=8.59179207911400411, type=longstraight;
sslxx: drift, l=0.024, type=longstraight; // skew sextupole
cplong: line=(hlxx, vlxx, qlxx, bpms, qlsxx, sxlxx, sslxx);
!!!!!!!!!!!!!!!! beginning of ring definition
fmagu01: fmag;
fmagd01: fmag;
dmagu01: dmag;
dmagd01: dmag;
cell01 : line = (sa, cpshort, sb, fmagu01, mins, dmagu01, dlong, cplong, drifte, dmagd01, mins, fmagd01, sc);
fmagu02: fmag;
fmagd02: fmag;
dmagu02: dmag;
dmagd02: dmag;
cell02 : line = (sa, cpshort, sb, fmagu02, mins, dmagu02, dlong, cplong, drifte, dmagd02, mins, fmagd02, sc);
fmagu03: fmag;
fmagd03: fmag;
dmagu03: dmag;
dmagd03: dmag;
cell03 : line = (sa, cpshort, sb, fmagu03, mins, dmagu03, dlong, cplong, drifte, dmagd03, mins, fmagd03, sc);
fmagu04: fmag;
fmagd04: fmag;
dmagu04: dmag;
dmagd04: dmag;
cell04 : line = (sa, cpshort, sb, fmagu04, mins, dmagu04, dlong, cplong, drifte, dmagd04, mins, fmagd04, sc);
fmagu05: fmag;
fmagd05: fmag;
dmagu05: dmag;
dmagd05: dmag;
cell05 : line = (sa, cpshort, sb, fmagu05, mins, dmagu05, dlong, cplong, drifte, dmagd05, mins, fmagd05, sc);
fmagu06: fmag;
fmagd06: fmag;
dmagu06: dmag;
dmagd06: dmag;
cell06 : line = (sa, cpshort, sb, fmagu06, mins, dmagu06, dlong, cplong, drifte, dmagd06, mins, fmagd06, sc);
fmagu07: fmag;
fmagd07: fmag;
dmagu07: dmag;
dmagd07: dmag;
cell07 : line = (sa, cpshort, sb, fmagu07, mins, dmagu07, dlong, cplong, drifte, dmagd07, mins, fmagd07, sc);
fmagu08: fmag;
fmagd08: fmag;
dmagu08: dmag;
dmagd08: dmag;
cell08 : line = (sa, cpshort, sb, fmagu08, mins, dmagu08, dlong, cplong, drifte, dmagd08, mins, fmagd08, sc);
fmagu09: fmag;
fmagd09: fmag;
dmagu09: dmag;
dmagd09: dmag;
cell09 : line = (sa, cpshort, sb, fmagu09, mins, dmagu09, dlong, cplong, drifte, dmagd09, mins, fmagd09, sc);
fmagu10: fmag;
fmagd10: fmag;
dmagu10: dmag;
dmagd10: dmag;
cell10 : line = (sa, cpshort, sb, fmagu10, mins, dmagu10, dlong, cplong, drifte, dmagd10, mins, fmagd10, sc);
fmagu11: fmag;
fmagd11: fmag;
dmagu11: dmag;
dmagd11: dmag;
cell11 : line = (sa, cpshort, sb, fmagu11, mins, dmagu11, dlong, cplong, drifte, dmagd11, mins, fmagd11, sc);
fmagu12: fmag;
fmagd12: fmag;
dmagu12: dmag;
dmagd12: dmag;
cell12 : line = (sa, cpshort, sb, fmagu12, mins, dmagu12, dlong, cplong, drifte, dmagd12, mins, fmagd12, sc);
fmagu13: fmag;
fmagd13: fmag;
dmagu13: dmag;
dmagd13: dmag;
cell13 : line = (sa, cpshort, sb, fmagu13, mins, dmagu13, dlong, cplong, drifte, dmagd13, mins, fmagd13, sc);
fmagu14: fmag;
fmagd14: fmag;
dmagu14: dmag;
dmagd14: dmag;
rf01: line=(drrf, rfc, drrf);
rf02: line=(drrf, rfc, drrf);
cell14: line = (sa, cpshort, sb, fmagu14, mins, dmagu14, drifta, rf01, driftb, rf02, dmidls, cplong, drifte, dmagd14, mins, fmagd14, sc);
fmagu15: fmag;
fmagd15: fmag;
dmagu15: dmag;
dmagd15: dmag;
rf03: line=(drrf, rfc, drrf);
rf04: line=(drrf, rfc, drrf);
cell15: line = (sa, cpshort, sb, fmagu15, mins, dmagu15, drifta, rf03, driftb, rf04, dmidls, cplong, drifte, dmagd15, mins, fmagd15, sc);
fmagu16: fmag;
fmagd16: fmag;
dmagu16: dmag;
dmagd16: dmag;
rf05: line=(drrf, rfc, drrf);
rf06: line=(drrf, rfc, drrf);
cell16: line = (sa, cpshort, sb, fmagu16, mins, dmagu16, drifta, rf05, driftb, rf06, dmidls, cplong, drifte, dmagd16, mins, fmagd16, sc);
fmagu17: fmag;
fmagd17: fmag;
dmagu17: dmag;
dmagd17: dmag;
rf07: line=(drrf, rfc, drrf);
rf08: line=(drrf, rfc, drrf);
cell17: line = (sa, cpshort, sb, fmagu17, mins, dmagu17, drifta, rf07, driftb, rf08, dmidls, cplong, drifte, dmagd17, mins, fmagd17, sc);
fmagu18: fmag;
fmagd18: fmag;
dmagu18: dmag;
dmagd18: dmag;
rf09: line=(drrf, rfc, drrf);
rf10: line=(drrf, rfc, drrf);
cell18: line = (sa, cpshort, sb, fmagu18, mins, dmagu18, drifta, rf09, driftb, rf10, dmidls, cplong, drifte, dmagd18, mins, fmagd18, sc);
fmagu19: fmag;
fmagd19: fmag;
dmagu19: dmag;
dmagd19: dmag;
rf11: line=(drrf, rfc, drrf);
rf12: line=(drrf, rfc, drrf);
cell19: line = (sa, cpshort, sb, fmagu19, mins, dmagu19, drifta, rf11, driftb, rf12, dmidls, cplong, drifte, dmagd19, mins, fmagd19, sc);
fmagu20: fmag;
fmagd20: fmag;
dmagu20: dmag;
dmagd20: dmag;
rf13: line=(drrf, rfc, drrf);
rf14: line=(drrf, rfc, drrf);
cell20: line = (sa, cpshort, sb, fmagu20, mins, dmagu20, drifta, rf13, driftb, rf14, dmidls, cplong, drifte, dmagd20, mins, fmagd20, sc);
fmagu21: fmag;
fmagd21: fmag;
dmagu21: dmag;
dmagd21: dmag;
rf15: line=(drrf, rfc, drrf);
rf16: line=(drrf, rfc, drrf);
cell21: line = (sa, cpshort, sb, fmagu21, mins, dmagu21, drifta, rf15, driftb, rf16, dmidls, cplong, drifte, dmagd21, mins, fmagd21, sc);
fmagu22: fmag;
fmagd22: fmag;
dmagu22: dmag;
dmagd22: dmag;
rf17: line=(drrf, rfc, drrf);
rf18: line=(drrf, rfc, drrf);
cell22: line = (sa, cpshort, sb, fmagu22, mins, dmagu22, drifta, rf17, driftb, rf18, dmidls, cplong, drifte, dmagd22, mins, fmagd22, sc);
fmagu23: fmag;
fmagd23: fmag;
dmagu23: dmag;
dmagd23: dmag;
rf19: line=(drrf, rfc, drrf);
rf20: line=(drrf, rfc, drrf);
cell23: line = (sa, cpshort, sb, fmagu23, mins, dmagu23, drifta, rf19, driftb, rf20, dmidls, cplong, drifte, dmagd23, mins, fmagd23, sc);
fmagu24: fmag;
fmagd24: fmag;
dmagu24: dmag;
dmagd24: dmag;
rf21: line=(drrf, rfc, drrf);
rf22: line=(drrf, rfc, drrf);
cell24: line = (sa, cpshort, sb, fmagu24, mins, dmagu24, drifta, rf21, driftb, rf22, dmidls, cplong, drifte, dmagd24, mins, fmagd24, sc);
booster: line=(cell01, cell02, cell03, cell04, cell05, cell06, cell07, cell08, cell09, cell10, cell11, cell12, cell13, cell14, cell15, cell16, cell17, cell18, cell19, cell20, cell21, cell22, cell23, cell24);
beam, particle=proton, energy=0.8+pmass;
!beam, particle=proton, energy=1.7386206689709858;
!beam, particle=proton, energy=0.8003486229709857+pmass;
Analyze
We run the following script to analyze correctness:
Script analysis_booster_simple.py
#!/usr/bin/env python3
#
# Copyright 2022-2026 ImpactX contributors
# Authors: Eric G. Stern, Axel Huebl, Chad Mitchell
# License: BSD-3-Clause-LBNL
#
# -*- coding: utf-8 -*-
import glob
import math
import re
import openpmd_api as io
import pandas as pd
from booster_impactx_lattice import get_lattice
from impactx import elements
def read_file(file_pattern):
for filename in glob.glob(file_pattern):
df = pd.read_csv(filename, delimiter=r"\s+")
if "step" not in df.columns:
step = int(re.findall(r"[0-9]+", filename)[0])
df["step"] = step
else:
df = df[df["step"] != "step"]
df = df.apply(pd.to_numeric)
yield df
def read_time_series(file_pattern):
"""Read in all CSV files from each MPI rank (and potentially OpenMP
thread). Concatenate into one Pandas dataframe.
Returns
-------
pandas.DataFrame
"""
return pd.concat(
read_file(file_pattern),
axis=0,
ignore_index=True,
) # .set_index('id')
# load the Booster lattice
lattice = elements.KnownElementsList()
lattice.extend(get_lattice())
# total lattice length (nominal Fermilab Booster circumference: 474.20 m)
total_s = sum(element.ds for element in lattice)
expected_s = 474.20
# tolerance loose enough to hold in single precision accumulation across the ring
assert math.isclose(total_s, expected_s, rel_tol=1.0e-4), (
f"lattice length {total_s} m does not match expected {expected_s} m"
)
# initial/final beam (TODO)
# series = io.Series("diags/openPMD/monitor.h5", io.Access.read_only)
# last_step = list(series.iterations)[-1]
# initial = series.iterations[1].particles["beam"].to_df()
# final = series.iterations[last_step].particles["beam"].to_df()
# compare number of particles (TODO)
# num_particles = 10000
# assert num_particles == len(initial)
# assert num_particles == len(final)
# TODO for Eric: add more tests as needed, e.g., checking beam moments
series = io.Series("diags/openPMD/monitor.h5", io.Access.read_only)
first_step = list(series.iterations)[0]
initial = series.iterations[first_step].particles["beam"].to_df()
last_step = list(series.iterations)[-1]
final = series.iterations[last_step].particles["beam"].to_df()
beta_ref = series.iterations[first_step].particles["beam"].get_attribute("beta_ref")
rbc = read_time_series("diags/reduced_beam_characteristics.*")
sigma_x = rbc["sigma_x"]
sigma_y = rbc["sigma_y"]
sigma_t = rbc["sigma_t"]
sigma_px = rbc["sigma_px"]
sigma_py = rbc["sigma_py"]
sigma_pt = rbc["sigma_pt"]
emittance_x = rbc["emittance_x"]
emittance_y = rbc["emittance_y"]
emittance_t = rbc["emittance_t"]
print("\t\tInitial\t\tFinal")
print("\t\t=======\t\t=====")
print(f"std x\t\t{sigma_x.iloc[0]:7g}\t{sigma_x.iloc[-1]:7g}")
print(f"std px\t\t{sigma_px.iloc[0]:7g}\t{sigma_px.iloc[-1]:7g}")
print(f"emittance x\t{emittance_x.iloc[0]:7g}\t{emittance_x.iloc[-1]:7g}")
print()
print(f"std y\t\t{sigma_y.iloc[0]:7g}\t{sigma_y.iloc[-1]:7g}")
print(f"std py\t\t{sigma_py.iloc[0]:7g}\t{sigma_py.iloc[-1]:7g}")
print(f"emittance y\t{emittance_y.iloc[0]:7g}\t{emittance_y.iloc[-1]:7g}")
print()
print(f"std t\t\t{sigma_t.iloc[0]:7g}\t\t{sigma_t.iloc[-1]:7g}")
print(f"std pt\t\t{sigma_pt.iloc[0]:7g}\t{sigma_pt.iloc[-1]:7g}")
print(f"emittance t\t{emittance_t.iloc[0]:7g}\t{emittance_t.iloc[-1]:7g}")
stdx_0 = 0.00836264
stdpx_0 = 0.000224866
stdy_0 = 0.00299539
stdpy_0 = 0.00057078
stdt_0 = 1.1694
stdpt_0 = 0.000927767
emitx_0 = 1.8803e-06
emity_0 = 1.70969e-06
emitt_0 = 0.00108493
assert math.isclose(sigma_x.iloc[0], stdx_0, rel_tol=2.0e-2)
assert math.isclose(sigma_x.iloc[-1], stdx_0, rel_tol=2.0e-2), (
"sigma_x change over one turn too large"
)
assert math.isclose(sigma_px.iloc[0], stdpx_0, rel_tol=2.0e-2)
assert math.isclose(sigma_px.iloc[-1], stdpx_0, rel_tol=2.0e-2), (
"sigma_px change over one turn too large"
)
assert math.isclose(emittance_x.iloc[0], emitx_0, rel_tol=2.0e-2)
assert math.isclose(emittance_x.iloc[1], emitx_0, rel_tol=2.0e-2), (
"emittance_x change over one turn too large"
)
assert math.isclose(sigma_y.iloc[0], stdy_0, rel_tol=2.0e-2)
assert math.isclose(sigma_y.iloc[-1], stdy_0, rel_tol=2.0e-2), (
"sigma_y change over one turn too large"
)
assert math.isclose(sigma_py.iloc[0], stdpy_0, rel_tol=2.0e-2)
assert math.isclose(sigma_py.iloc[-1], stdpy_0, rel_tol=2.0e-2), (
"sigma_py change over one turn too large"
)
assert math.isclose(emittance_y.iloc[0], emity_0, rel_tol=2.0e-2)
assert math.isclose(emittance_y.iloc[1], emity_0, rel_tol=2.0e-2), (
"emittance_y change over one turn too large"
)
# longitudinal distribution is not matched and executes
# synchrotron dipole and quadrupole oscillations
The second moments of the transverse particle distribution after the FODO cell should coincide with the second moments of the particle distribution before the FODO cell, to within the level expected due to noise due to statistical sampling.
Visualize
You can run the following script to visualize the beam evolution over time:
Script plot_simple_booster.py
#!/usr/bin/env python3
#
# Copyright 2022-2026 ImpactX contributors
# Authors: Eric G. Stern, Axel Huebl, Chad Mitchell
# License: BSD-3-Clause-LBNL
#
# -*- coding: utf-8 -*-
import argparse
import glob
import re
import matplotlib.pyplot as plt
import openpmd_api as io
import pandas as pd
import scipy
def read_file(file_pattern):
for filename in glob.glob(file_pattern):
df = pd.read_csv(filename, delimiter=r"\s+")
if "step" not in df.columns:
step = int(re.findall(r"[0-9]+", filename)[0])
df["step"] = step
else:
df = df[df["step"] != "step"]
df = df.apply(pd.to_numeric)
yield df
def read_time_series(file_pattern):
"""Read in all CSV files from each MPI rank (and potentially OpenMP
thread). Concatenate into one Pandas dataframe.
Returns
-------
pandas.DataFrame
"""
return pd.concat(
read_file(file_pattern),
axis=0,
ignore_index=True,
) # .set_index('id')
# options to run this script
parser = argparse.ArgumentParser(description="Plot the FODO benchmark.")
parser.add_argument(
"--save-png", action="store_true", help="non-interactive run: save to PNGs"
)
args = parser.parse_args()
series = io.Series("diags/openPMD/monitor.h5", io.Access.read_only)
first_step = list(series.iterations)[0]
initial = series.iterations[first_step].particles["beam"].to_df()
last_step = list(series.iterations)[-1]
final = series.iterations[last_step].particles["beam"].to_df()
beta_ref = series.iterations[first_step].particles["beam"].get_attribute("beta_ref")
# columns in rbc file
# step s mean_x min_x max_x mean_y min_y max_y mean_t min_t max_t sigma_x sigma_y sigma_t mean_px min_px max_px mean_py min_py max_py mean_pt min_pt max_pt sigma_px sigma_py sigma_pt emittance_x emittance_y emittance_t alpha_x alpha_y alpha_t beta_x beta_y beta_t dispersion_x dispersion_px dispersion_y dispersion_py emittance_xn emittance_yn emittance_tn charge_C
rbc = read_time_series("diags/reduced_beam_characteristics.*")
s = rbc["s"]
min_x = rbc["min_x"]
max_x = rbc["max_x"]
min_y = rbc["min_y"]
max_y = rbc["max_y"]
min_t = rbc["min_t"]
max_t = rbc["max_t"]
sigma_x = rbc["sigma_x"]
sigma_y = rbc["sigma_y"]
sigma_t = rbc["sigma_t"]
sigma_pt = rbc["sigma_pt"]
charge = rbc["charge_C"] / scipy.constants.eV
f, ax = plt.subplots(2, 2, sharex="row")
ax[0, 0].set_title(r"$\sigma_x$")
ax[0, 0].plot(s, sigma_x * 1000.0)
ax[0, 0].set_ylabel(r"$\sigma_x$ [mm]")
ax[0, 1].set_title(r"$\sigma_y$")
ax[0, 1].plot(s, sigma_y * 1000.0)
ax[0, 1].set_ylabel(r"$\sigma_y$ [mm]")
ax[1, 0].set_title(r"$\sigma_t$")
ax[1, 0].plot(s, sigma_t, label=r"$sigma_t$ [m]")
ax[1, 0].set_xlabel("s [m]")
ax[1, 0].set_ylabel(r"$\sigma_t$ [m]")
ax[1, 1].set_title(r"$\sigma_{pt}$")
ax[1, 1].plot(s, sigma_pt * 1000.0, label="sigma_pt")
ax[1, 1].set_ylabel(r"$\sigma_{pt} \, [\times 1000]$")
ax[1, 1].set_xlabel("s [m]")
plt.tight_layout()
if args.save_png:
plt.savefig("simple_booster_sigma.png")
else:
plt.show()
f, ax = plt.subplots(2, 2, sharex="row", sharey="row")
ax[0, 0].set_title("x vs. px initial")
ax[0, 0].plot(
initial["position_x"] * 1000,
initial["momentum_x"] * 1000,
".",
label="initial x vs. px",
)
ax[0, 0].set_xlabel("x [mm]")
ax[0, 0].set_ylabel("px [x1000]")
ax[0, 1].set_title("x vs. px final")
ax[0, 1].plot(
final["position_x"] * 1000,
final["momentum_x"] * 1000,
".",
label="final x vs. px",
)
ax[0, 1].set_xlabel("x [mm]")
ax[1, 0].set_title("y vs. py initial")
ax[1, 0].plot(
initial["position_y"] * 1000,
initial["momentum_y"] * 1000,
".",
label="initial y vs. py",
)
ax[1, 0].set_xlabel("y [mm]")
ax[1, 0].set_ylabel("py [x1000]")
# ax[1, 0].legend(loc="best")
ax[1, 1].set_title("y vs. py final")
ax[1, 1].plot(
final["position_y"] * 1000,
final["momentum_y"] * 1000,
".",
label="final y vs. py",
)
ax[1, 1].set_xlabel("y [mm]")
plt.tight_layout()
if args.save_png:
plt.savefig("simple_booster_scatter.png")
else:
plt.show()
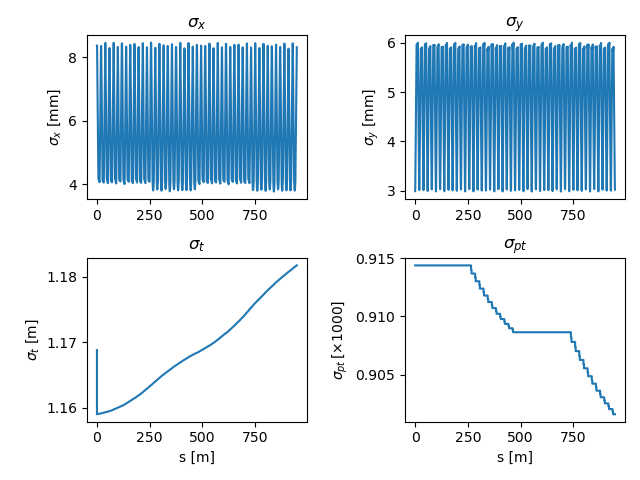
Fig. 14 Evolution of beam sigmas over two turns in the simple Booster model.
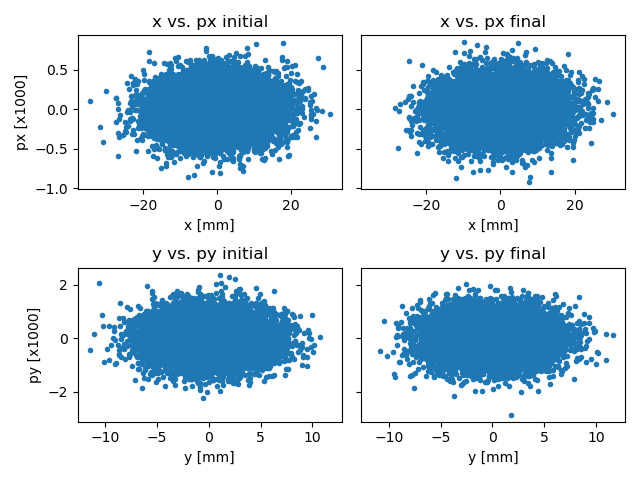
Fig. 15 Simple Booster initial and final phase space.
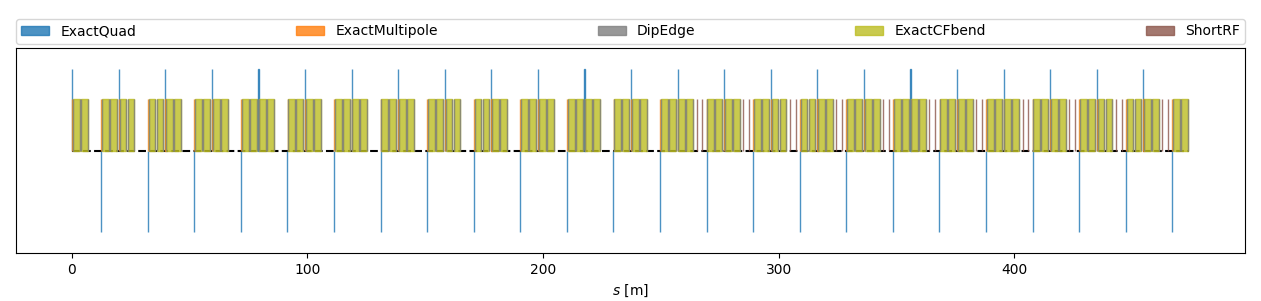
Lattice Survey
Generate a survey of the layout of the Simple Booster machine
Script plot_simple_booster_survey.py
#!/usr/bin/env python3
#
# Copyright 2022-2026 ImpactX contributors
# Authors: Eric G. Stern, Axel Huebl, Chad Mitchell
# License: BSD-3-Clause-LBNL
#
# -*- coding: utf-8 -*-
import argparse
import matplotlib.pyplot as plt
from booster_impactx_lattice import get_lattice
from impactx import ImpactX, elements
# options to run this script
parser = argparse.ArgumentParser(
description="Plot the Fermilab Booster lattice survey."
)
parser.add_argument(
"--save-png", action="store_true", help="non-interactive run: save to PNGs"
)
args = parser.parse_args()
# reference particle (800 MeV proton, as in run_simple_booster.py)
sim = ImpactX()
sim.init_grids()
ref = sim.particle_container().ref_particle()
ref.set_species("proton").set_kin_energy_MeV(800.0)
# load the Booster lattice
lattice = elements.KnownElementsList()
lattice.extend(get_lattice())
# survey plot
fig = plt.figure(figsize=(12, 4.8))
lattice.plot_survey(ref=ref)
plt.tight_layout()
if args.save_png:
plt.savefig("simple_booster_survey.png")
else:
plt.show()